Uni-Encoder Meets Multi-Encoders: Representation Before Fusion for Brain Tumor Segmentation with Missing Modalities
Abstract
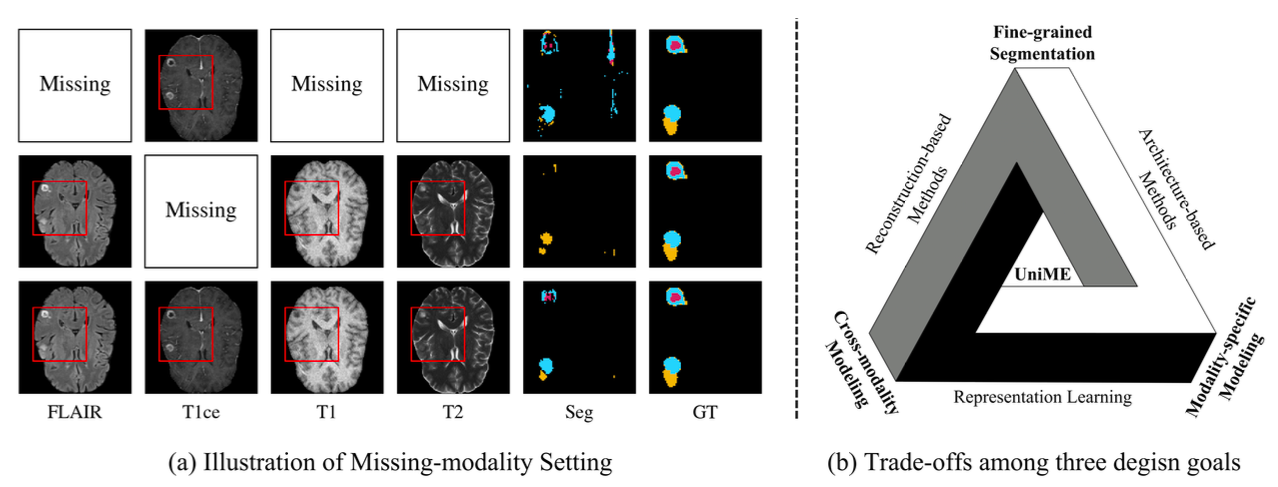
Multimodal MRI provides complementary information for brain tumor segmentation, but clinical scans often suffer from missing modalities, which significantly degrades performance. Existing methods typically struggle to balance three key objectives: capturing fine-grained anatomical structures, modeling cross-modal complementarity, and effectively exploiting available modalities.
We propose UniME (Uni-Encoder Meets Multi-Encoders), a two-stage heterogeneous framework that explicitly decouples representation learning from segmentation. In Stage 1, a unified ViT-based encoder is pretrained using masked self-supervision to learn modality-agnostic representations robust to missing modalities. In Stage 2, modality-specific CNN encoders are introduced to extract high-resolution, multi-scale features, which are fused with the global representation for accurate segmentation.
This "representation before fusion" paradigm enables UniME to simultaneously model global cross-modal semantics and fine-grained local structures. Extensive experiments on BraTS 2023 and BraTS 2024 demonstrate that UniME consistently outperforms existing state-of-the-art methods under incomplete modality settings.
Key Contributions
- We propose UniME, a two-stage heterogeneous framework that decouples representation learning from segmentation, addressing the fundamental trade-off in missing-modality segmentation.
- We introduce a unified ViT-based Uni-Encoder pretrained with masked self-supervision, enabling robust modality-agnostic representation learning under incomplete inputs.
- We design modality-specific Multi-Encoders to capture high-resolution, multi-scale features, complementing the global representation with fine-grained structural details.
- We establish a "representation before fusion" paradigm, which effectively integrates global semantic understanding and local structural modeling for improved segmentation performance.
Motivation
Brain tumor segmentation in clinical practice faces a fundamental challenge: complete sets of MRI modalities are often unavailable. Existing methods address this by either synthesizing missing modalities, using knowledge distillation, or redesigning network architectures — but each approach involves trade-offs that limit overall performance.
Method Overview
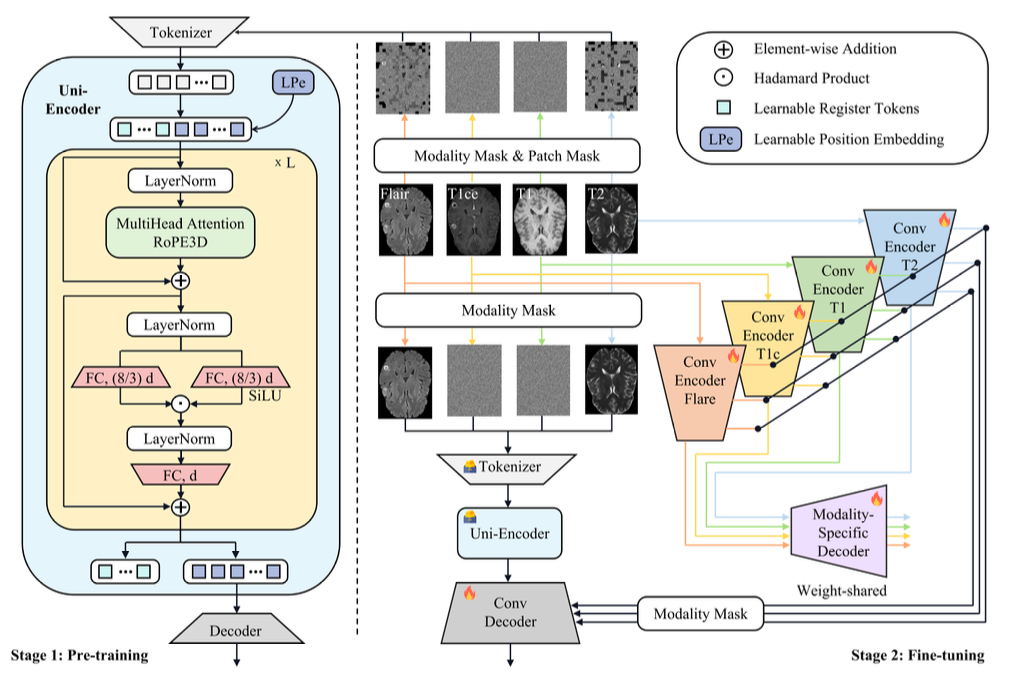
UniME follows a two-stage heterogeneous design that separates representation learning from segmentation.
In Stage 1, a single ViT-based Uni-Encoder is pretrained using masked self-supervised learning. By combining modality-level and patch-level masking, the encoder learns a unified representation that is robust to missing modalities and captures global cross-modal semantics.
In Stage 2, modality-specific CNN Multi-Encoders are introduced to extract high-resolution, multi-scale features from available modalities. These fine-grained features are fused with the global representation from the Uni-Encoder to produce accurate segmentation results.
This design follows a "representation before fusion" principle: instead of directly fusing modality-specific features, UniME first learns a unified representation and then enhances it with modality-specific details, effectively balancing global semantics and local structures.
Results
We evaluate UniME on BraTS 2023 and BraTS 2024 under various missing-modality settings. The proposed method consistently outperforms state-of-the-art approaches across different tumor regions (WT, TC, ET).
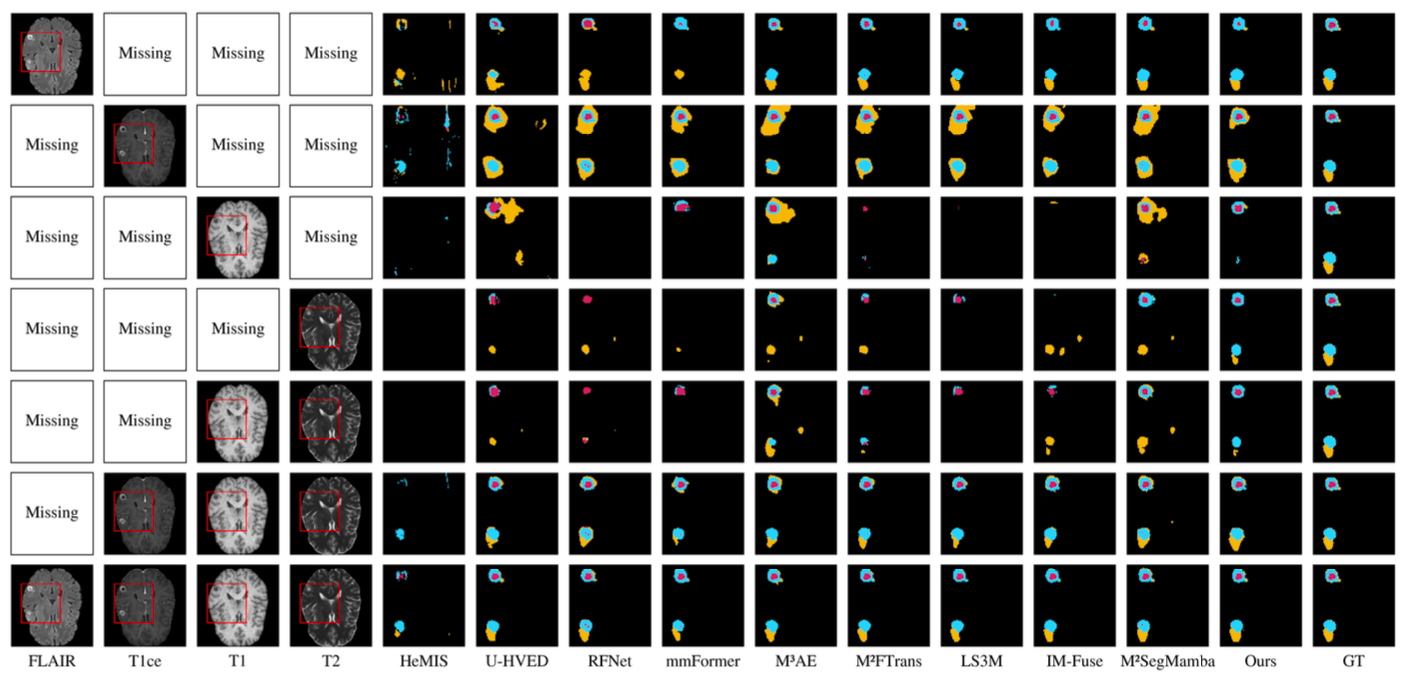

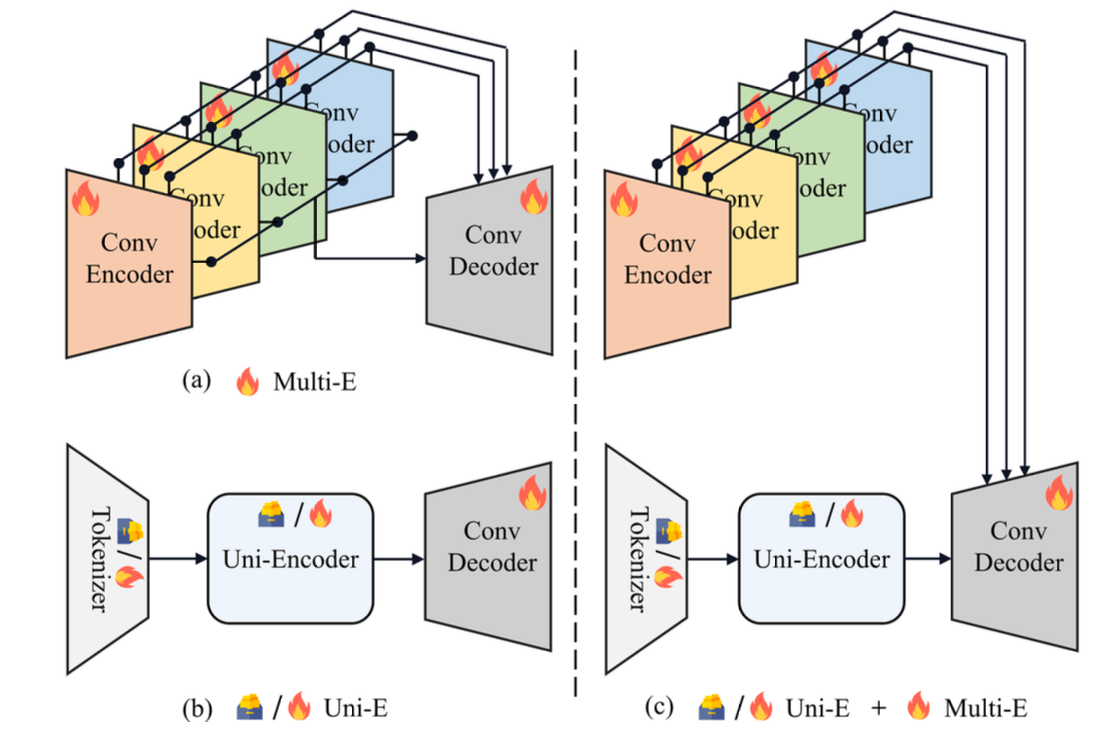
Figure 4 further validates the effectiveness of the two-stage design. The combination of Uni-Encoder and Multi-Encoders significantly improves performance compared to single-encoder or multi-encoder-only designs, highlighting their complementary roles.
Citation
@article{song2026unime,
title={Uni-Encoder Meets Multi-Encoders: Representation Before Fusion for Brain Tumor Segmentation with Missing Modalities},
author={Song, Peibo and Xue, Xiaotian and Zhang, Jinshuo and Wang, Zihao and Liu, Jinhua and Fu, Shujun and Bao, Fangxun and Yeo, Si Yong},
booktitle={Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR)},
year={2026}
}