Prototypical Representation Learning for Multi-Site Domain Generalization in Schizophrenia Diagnosis
Abstract
Schizophrenia diagnosis from multi-site resting-state fMRI is challenged by substantial site-induced distribution shifts, which often hinder the generalization of brain functional network (BFN)-based classifiers. Existing domain generalization methods usually assume that each class has consistent structures across sites, but this assumption is often violated by the inherent diversity within each class.
We propose a prototypical domain generalization framework for multi-site schizophrenia diagnosis. A transformer encoder first captures global inter-regional dependencies from BFNs to obtain discriminative subject-level embeddings. A site-independence module regularized by Hilbert-Schmidt Independence Criterion (HSIC) then encourages site-invariant feature learning. Projected features are softly assigned to multiple class-specific prototypes via the Sinkhorn-Knopp algorithm, with prototypes updated using exponential moving average (EMA). Maximum likelihood estimation (MLE) loss further refines feature-to-prototype matching probabilities, while prototype contrastive and alignment losses promote inter-class separation and intra-class compactness.
Experiments on FBIRN and BSNIP show that the proposed method achieves strong generalization performance on unseen sites, reaching 88.89%±2.22% and 86.05%±1.64% accuracy, respectively.
Key Contributions
- A multi-site domain generalization framework for schizophrenia diagnosis that combines transformer-based BFN representation learning with prototype learning.
- A site-independence module with HSIC regularization to suppress site-specific bias and encourage site-invariant embeddings.
- A multi-prototype learning strategy with Sinkhorn-based soft assignment and EMA prototype updates to better model intra-class diversity.
- A joint optimization objective combining classification, MLE, prototype contrastive, and prototype alignment losses to improve both separability and compactness.
- Identification of discriminative temporal brain regions associated with schizophrenia, providing biologically relevant interpretation.
Method Overview
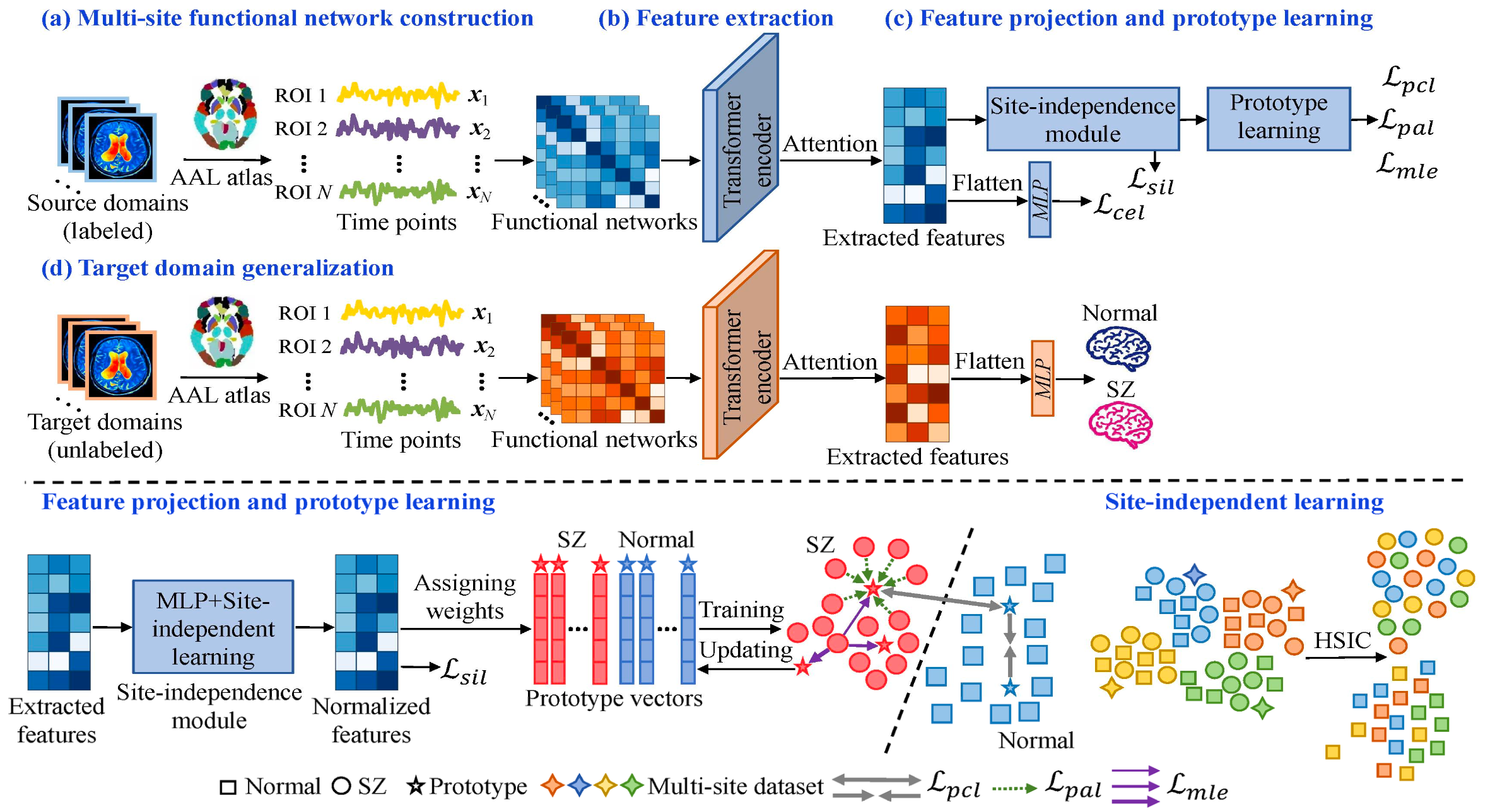
The proposed framework contains three major stages. First, BFNs constructed from resting-state fMRI are processed by a transformer encoder, which models global inter-regional dependencies and produces subject-level embeddings. Second, these features are passed through an MLP-based projection module regularized by HSIC to reduce dependence on site information and improve site invariance. Third, a prototype learning module assigns projected features to multiple class-specific prototypes using the Sinkhorn-Knopp algorithm. Prototypes are updated with EMA, while MLE, prototype contrastive, and prototype alignment losses jointly improve feature-to-prototype matching and class discrimination.
Results & Performance
The proposed framework was evaluated under a leave-one-site-out domain generalization protocol, where one site was held out as the unseen target domain and the remaining sites were used for training. Across two independent schizophrenia datasets, FBIRN and BSNIP, the method achieved strong and consistent performance against a broad range of comparison methods.
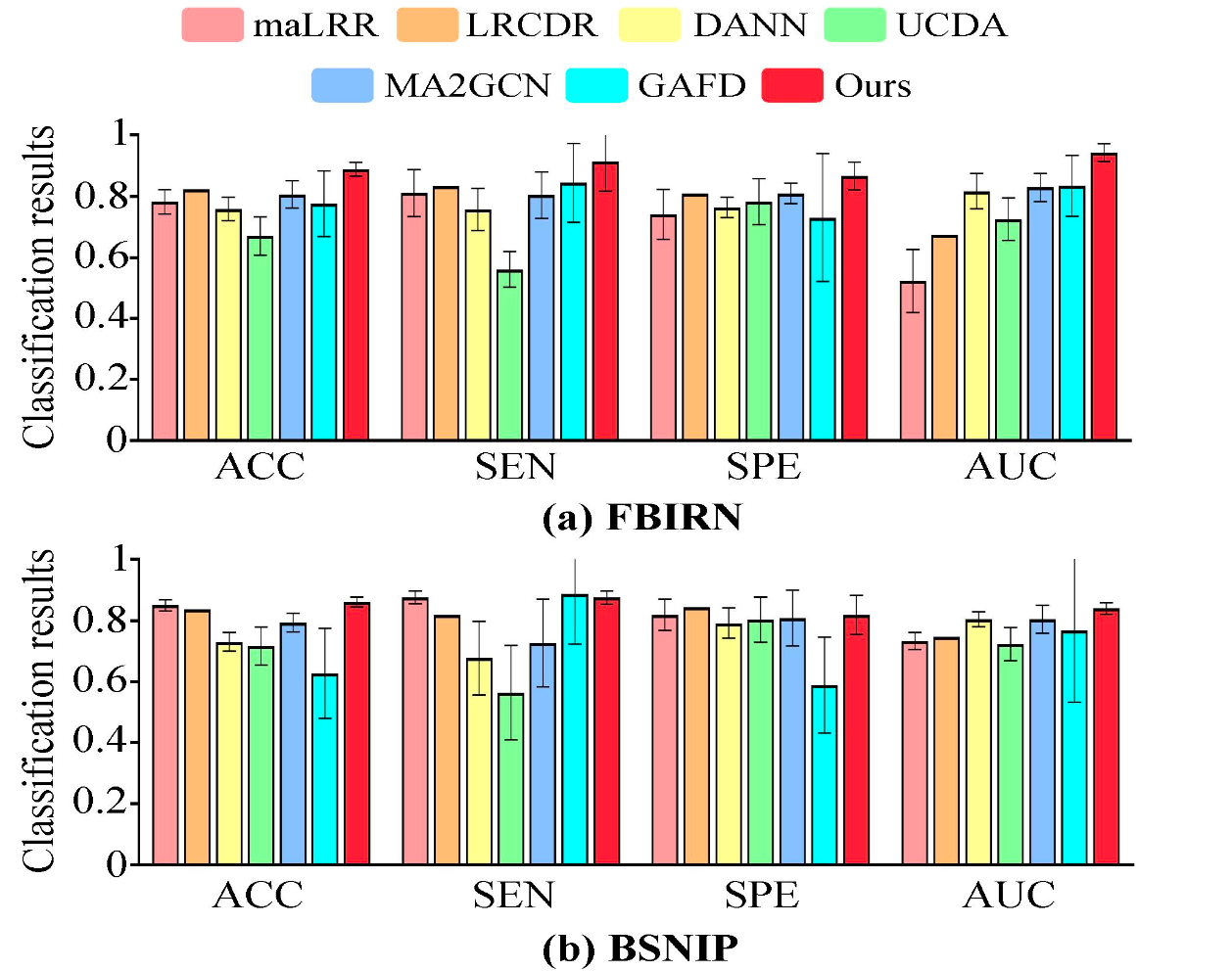
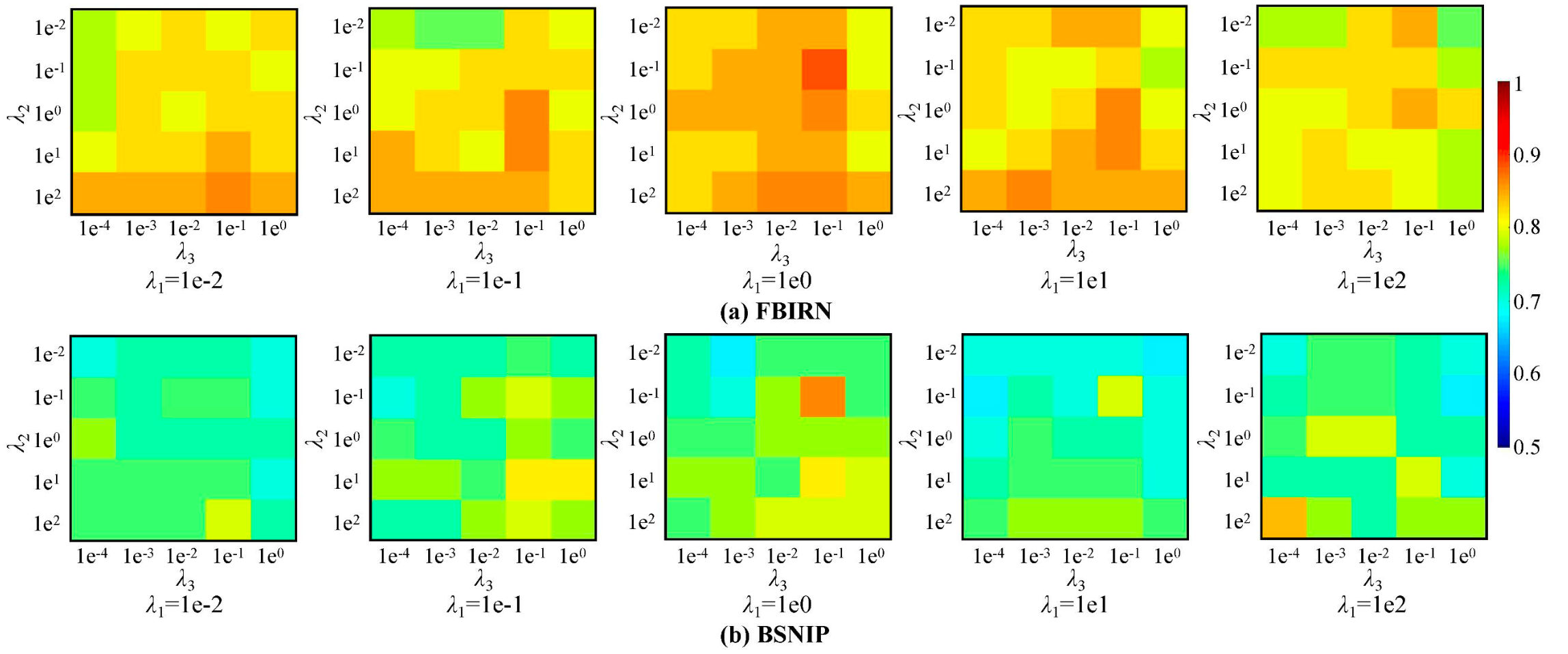
On the two benchmark datasets, the method achieved 88.89%±2.22% accuracy and 94.33%±2.95% AUC on FBIRN, and 86.05%±1.64% accuracy on BSNIP. The ablation study further showed that the MLE loss, contrastive loss, and alignment loss all contributed to the final performance, with MLE playing the most important role.
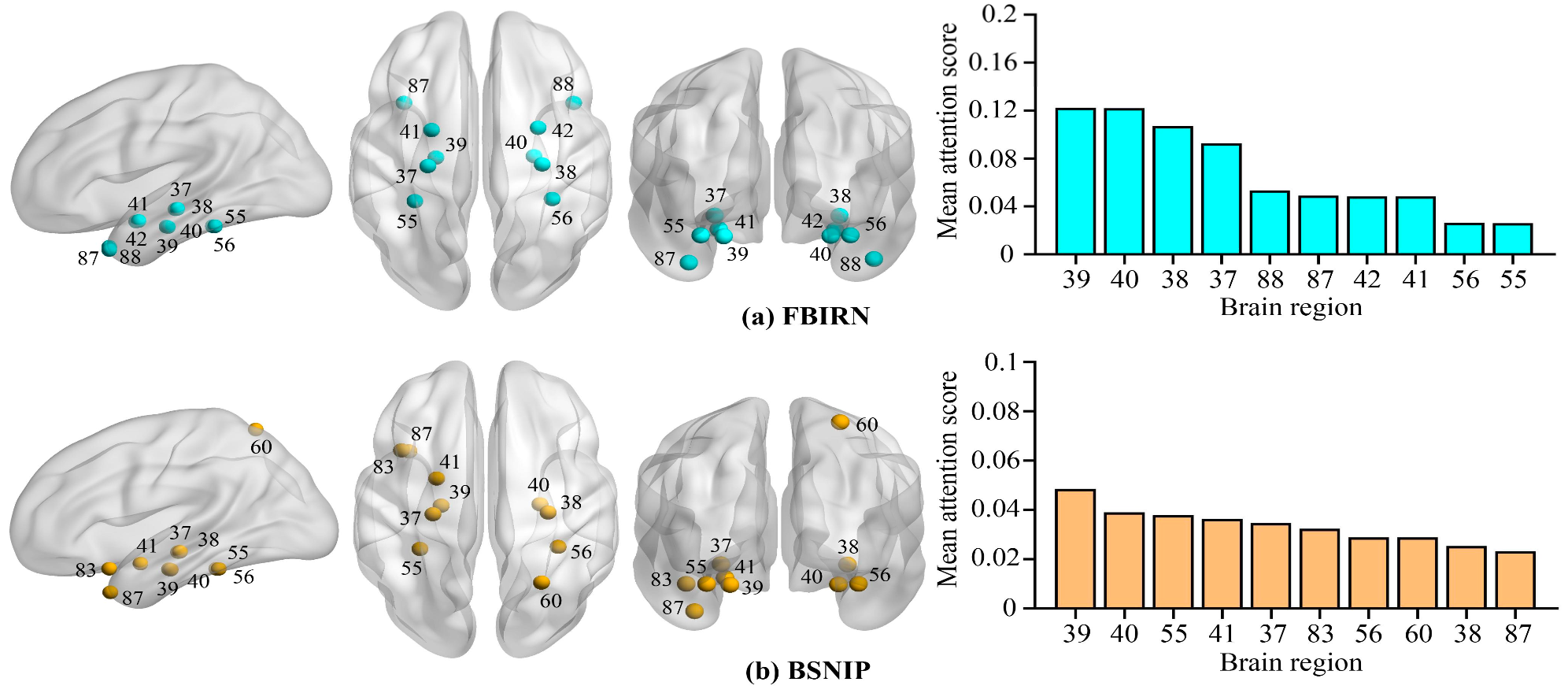
To provide neurological interpretability, the model identifies the top-10 most discriminative brain regions for schizophrenia classification. Temporal lobe regions were repeatedly highlighted across both datasets.
Citation
@article{ji2026prototypical,
title={Prototypical Representation Learning for Multi-Site Domain Generalization in Schizophrenia Diagnosis},
author={Ji, Yixin and Calhoun, Vince D and Zhang, Jin and Zhu, Qi and Li, Shengrong and Mathalon, Daniel H and Yeo, Si Yong and Zhang, Daoqiang and Qi, Shile},
journal={IEEE Transactions on Biomedical Engineering},
year={2026},
doi={10.1109/TBME.2026.3658874},
publisher={IEEE}
}