Exploiting Low-Dimensional Manifold of Features for Few-Shot Whole Slide Image Classification
Abstract
Few-shot Whole Slide Image (WSI) classification is severely hampered by overfitting. We argue that this is not merely a data-scarcity issue but a fundamentally geometric problem. Grounded in the manifold hypothesis, our analysis shows that features from pathology foundation models exhibit a low-dimensional manifold geometry that is easily perturbed by downstream models.
This reveals a key issue in downstream multiple instance learning models: linear layers are geometry-agnostic and can distort the manifold geometry of the features. To address this, we propose the Manifold Residual (MR) block, a plug-and-play module that is explicitly geometry-aware.
The MR block reframes the linear layer as residual learning and decouples it into two pathways: (1) a fixed random matrix serving as a geometric anchor that approximately preserves topology while also acting as a spectral shaper; and (2) a trainable low-rank residual pathway for task-specific adaptation, with its structural bottleneck explicitly mirroring the low effective rank of the features.
This decoupling imposes a structured inductive bias, reduces learning to a simpler residual fitting task, and achieves strong performance with significantly fewer parameters.
Motivation: Feature Geometry Analysis
A key insight of this work is that pathology foundation model features lie on a low-dimensional manifold. Standard linear layers distort this intrinsic geometry, causing overfitting in the few-shot regime. Our analysis provides both theoretical and empirical evidence for this observation.
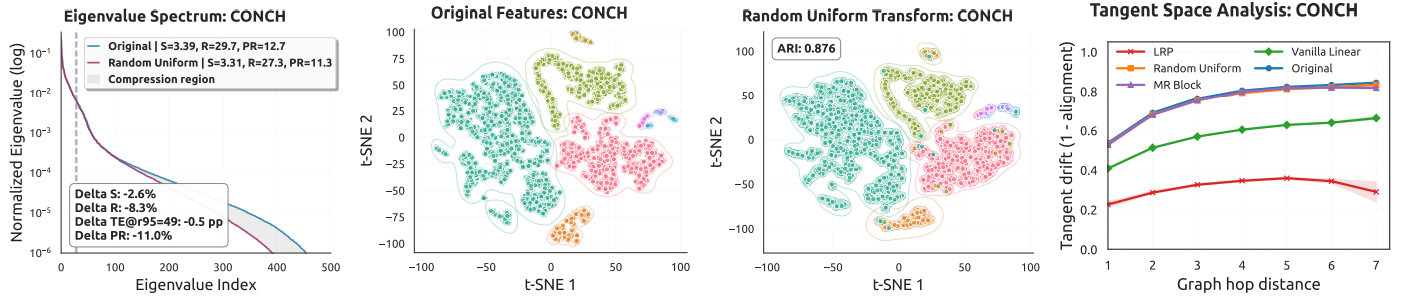
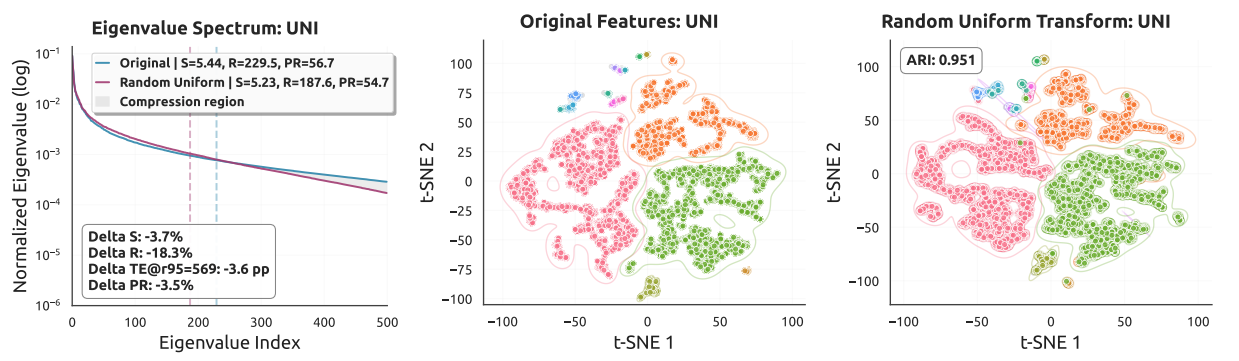
Key Contributions
- Geometric reframing: We show that pathology foundation model features lie on a low-dimensional manifold and that standard linear layers can disrupt this fragile geometry in few-shot settings.
- Manifold Residual (MR) block: We propose a theoretically grounded, plug-and-play replacement for internal linear layers that combines a geometric anchor with a low-rank residual pathway.
- Parameter efficiency: MR-enhanced models match or surpass much larger competing methods while using substantially fewer trainable parameters.
- Broad applicability: The MR block is backbone-agnostic and can be integrated into different multiple instance learning (MIL) frameworks.
Method Overview
The core contribution is the Manifold Residual (MR) Block, which reformulates a standard linear layer into two decoupled pathways. Rather than treating the transformation as a single learned mapping, the MR block separates geometric preservation from task-specific adaptation.
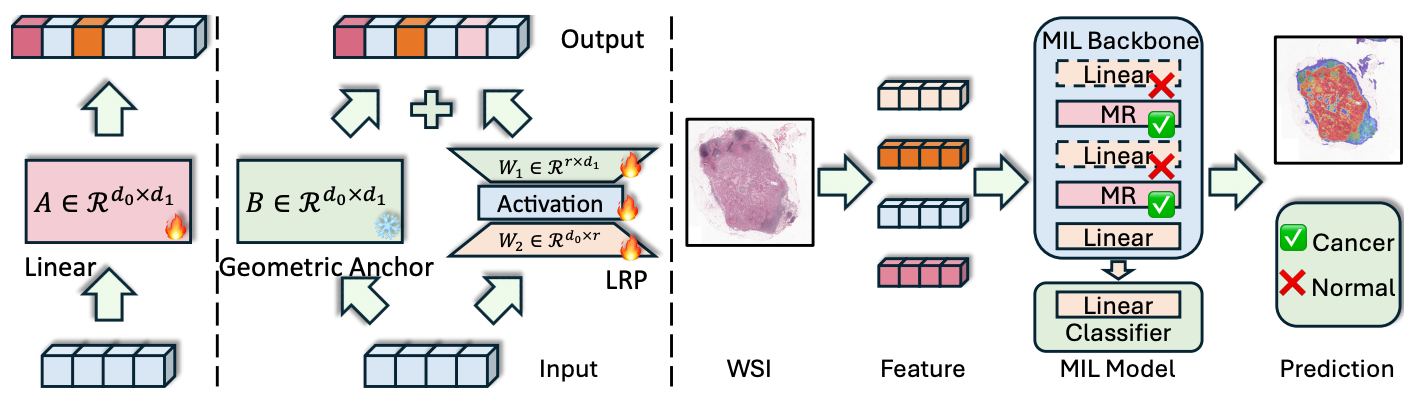
The MR block consists of two components:
- Geometric Anchor (fixed random matrix): Initialized with a random distribution and kept frozen during training. It approximately preserves the topology of the feature manifold and also acts as a spectral shaper.
- Low-Rank Residual Pathway (LRP): A trainable bottleneck with rank r ≪ min(d₀, d₁), explicitly reflecting the low effective rank of foundation model features. It learns task-specific adaptation as a residual on top of the geometric anchor.
This decoupling imposes a structured inductive bias and is particularly beneficial in the few-shot regime.
Results & Performance
MR-enhanced variants consistently improve over their corresponding baselines across multiple datasets, shot counts, and MIL backbones. The gains are strongest in the extreme few-shot regime and remain competitive at larger shot counts.
- Camelyon16, TCGA-NSCLC, and TCGA-RCC: MR-based models match or surpass strong recent methods while using far fewer trainable parameters.
- Boehmk and Trastuzumab: MR-ABMIL and MR-CATE achieve strong treatment-response prediction performance in challenging clinical settings.
- Parameter efficiency: Replacing vanilla linear layers with MR blocks reduces trainable parameters substantially while maintaining or improving performance.
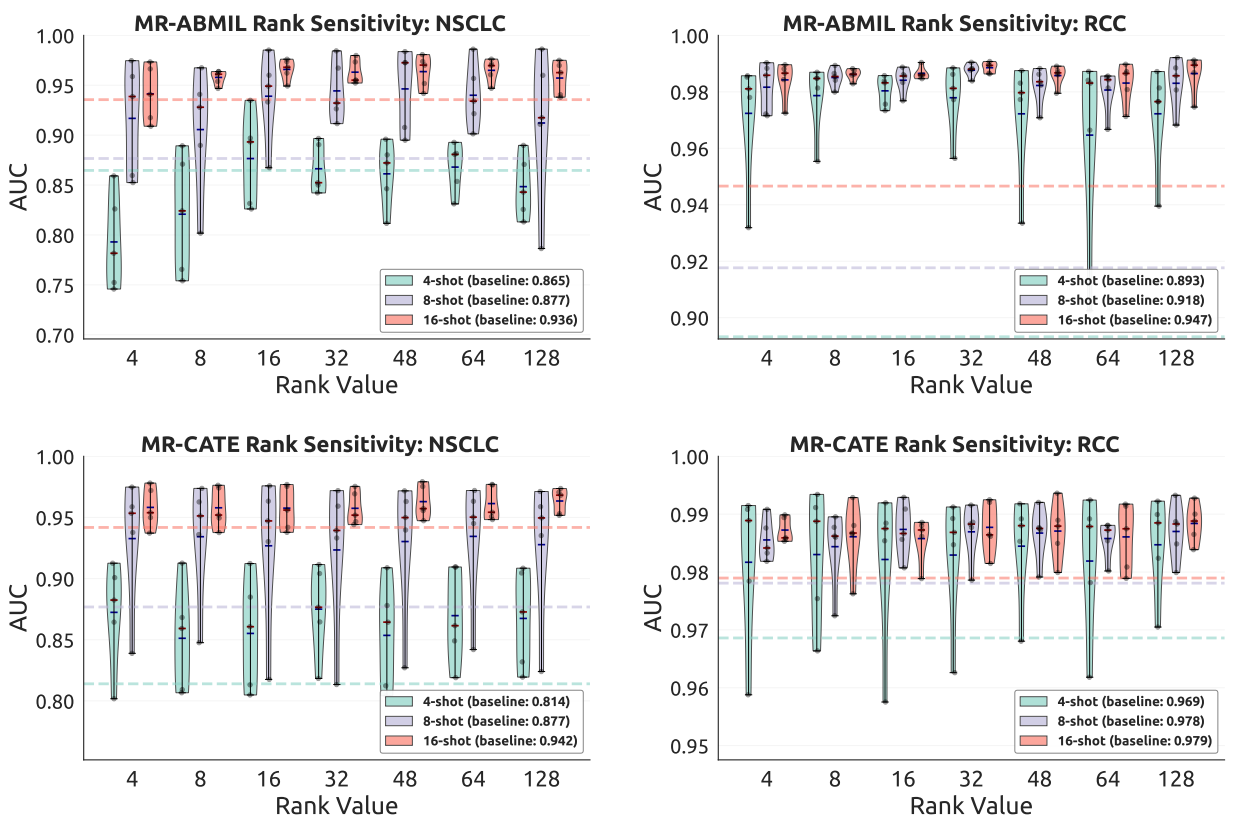
Figure 3 illustrates the sensitivity analysis of the bottleneck rank r, and the empirical trend supports the design principle of aligning the residual bottleneck with the intrinsic effective rank of the feature manifold.
Datasets
- Camelyon16: Tumor detection in breast lymph node whole slide images.
- TCGA-NSCLC: Non-small cell lung cancer subtyping from The Cancer Genome Atlas.
- TCGA-RCC: Renal cell carcinoma subtyping from The Cancer Genome Atlas.
- Boehmk: Treatment response prediction dataset.
- Trastuzumab: Treatment response prediction dataset.
Experiments are conducted across few-shot settings with k ∈ {2, 4, 8, 16, 32, 64} shots, using multiple pathology foundation model feature extractors including CONCH and UNI.
Citation
@inproceedings{xiong2026lmfs,
title={Exploiting Low-Dimensional Manifold of Features for Few-Shot Whole Slide Image Classification},
author={Xiong, Conghao and Guo, Zhengrui and Xu, Zhe and Zhang, Yifei and Tong, Raymond Kai-yu and Yeo, Si Yong and Chen, Hao and Sung, Joseph J. Y. and King, Irwin},
booktitle={International Conference on Learning Representations (ICLR)},
year={2026}
}
Accepted at ICLR 2026
International Conference on Learning Representations